|
aeruginosa data, we collected publicly available P. After validating a method for high-throughput read mapping of P. To support the exploration of public RNA-seq data and to further the development of resources that leverage these data, we present a computationally efficient method to reprocess RNA-seq data sets (see Fig. 1 for an overview). aeruginosa expression profiles from microarray and RNA sequencing (RNA-seq) technologies are publicly available. aeruginosa research is reflected in the abundance of transcriptional data sets in public databases, including those hosted by the National Center for Biotechnology Information (NCBI), such as the Sequence Read Archive (SRA) and the Gene Expression Omnibus (GEO), and those hosted by the European Molecular Biology Laboratory European Bioinformatics Institute (EMBL-EBI), such as the European Nucleotide Archive (ENA). aeruginosa studies use the laboratory strains PAO1 and PA14 as models for the study of transcriptional regulation, and many studies have also examined gene expression in clinical isolates. aeruginosa genome ( 7 – 9), transcriptional profiling across conditions and mutant genotypes has been a fruitful approach to better understand P. Given the unusually high numbers of transcription factors, sigma factors, and two-component systems in the P. aeruginosa is partly attributable to adaptive behavioral changes driven by gene expression. aeruginosa is also found in soil ( 2) and freshwater ( 3, 4), and it is cultured for biotechnology applications ( 5, 6). The opportunistic pathogen Pseudomonas aeruginosa causes infections in many body sites and is commonly found in chronic lung infections of people with cystic fibrosis ( 1), where it is difficult to eradicate, and the factors that lead to persistence are not fully understood. 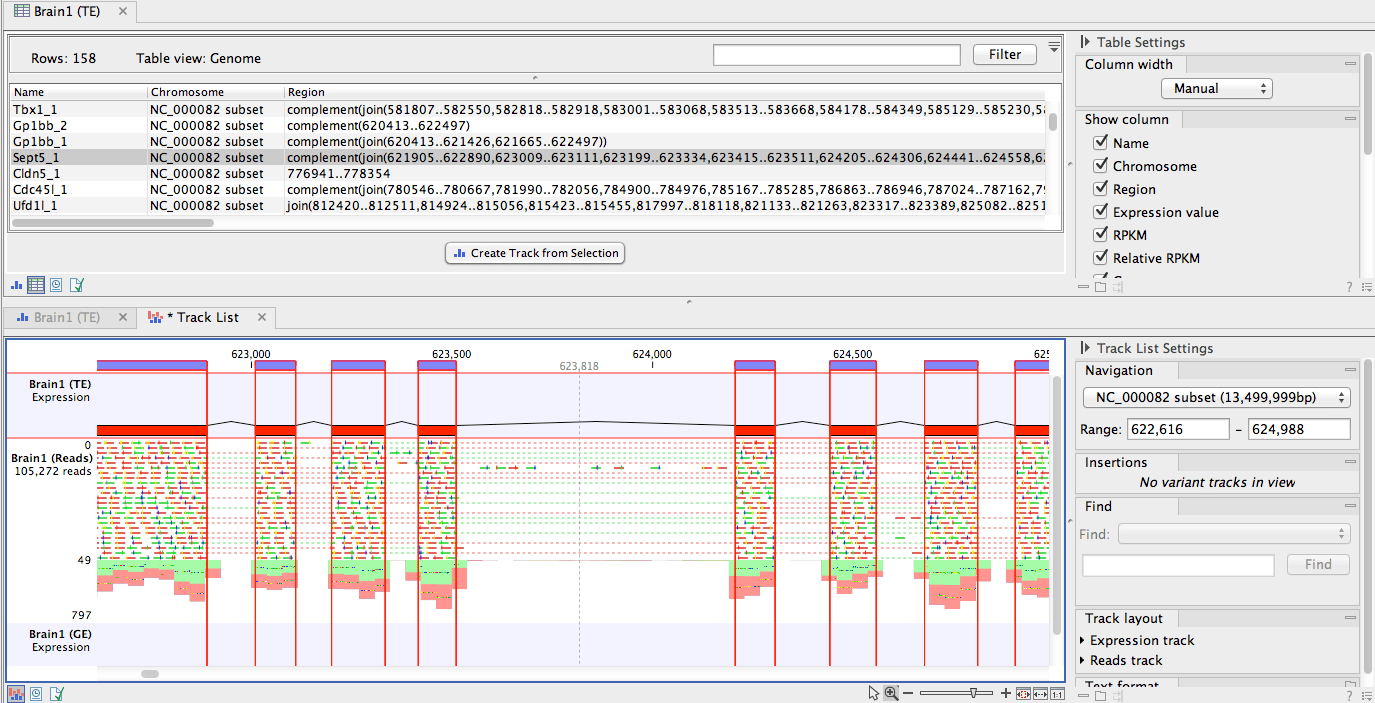
aeruginosa RNA-seq data to be leveraged for diverse research goals. Our processing and quality control methods provide a scalable framework for taking advantage of the troves of biological information hibernating in the depths of microbial gene expression data and yield useful tools for P. Finally, we developed an algorithm to incorporate new data as they are deposited into the SRA. Since the RNA-seq data were generated using diverse strains, we report the effects of mapping samples to noncognate reference genomes by separately analyzing all samples mapped to cDNA reference genomes for strains PAO1 and PA14, two divergent strains that were used to generate most of the samples. The filtering and normalization steps greatly improved gene expression correlations for genes within the same operon or regulon across the 2,333 samples. We developed filtering criteria to exclude samples with aberrant levels of housekeeping gene expression or an unexpected number of genes with no reported values and normalized the filtered compendia using the ratio-of-medians method. Raw sequence data were uniformly processed using the Salmon pseudoaligner, and this read mapping method was validated by comparison to a direct alignment method. In this work, the transcriptional profiles from hundreds of studies performed by over 75 research groups were reanalyzed in aggregate to create a powerful tool for hypothesis generation and testing. Error probabilities.Thousands of Pseudomonas aeruginosa RNA sequencing (RNA-seq) gene expression profiles are publicly available via the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA). Ewing B, Green P (1998) Base-calling of automated sequencer traces using phred.Koch CM, Chiu SF, Akbarpour M, Bharat A, Ridge KM, Bartom ET, Winter DR (2018) A Beginner’s guide to analysis of RNA sequencing data. 
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-seq.Byron SA, Van Keuren-Jensen KR, Engelthaler DM, Carpten JD, Craig DW (2016) Translating RNA sequencing into clinical diagnostics: opportunities and challenges.Ozsolak F, Milos PM (2011) RNA sequencing: advances, challenges and opportunities.Royce TE, Rozowsky JS, Gerstein MB (2007) Toward a universal microarray: prediction of gene expression through nearest-neighbor probe sequence identification.Okoniewski MJ, Miller CJ (2006) Hybridization interactions between probesets in short oligo microarrays lead to spurious correlations.van Hal NL, Vorst O, van Houwelingen AM, Kok EJ, Peijnenburg A, Aharoni A, van Tunen AJ, Keijer J (2000) The application of DNA microarrays in gene expression analysis.Wang Z, Gerstein M, Snyder M (2009) RNA-seq: a revolutionary tool for transcriptomics.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
